Cultivars, strains, chemovars (whatever you love to call them) that were sequenced with our StrainSEEK® Version 2 technology have been mapped to the new Jamaican Lion reference and their reports have been updated on Kannapedia.net.
StrainSEEK® customers who visit their report pages will find a new data visualization showing Bt:Bd allele coverage, changes to the Genetic Relatives and Most Genetically Distant Strains tables, new VCF files, and a new blockchain registration.
To create the Jamaican Lion assembly, over 180 billion bases were sequenced with Pacific Biosciences latest Version 6 Chemistry, allowing us to select reads that average 60,000 base pairs in length with 15x coverage. This is an order of magnitude more contiguous than anything produced to date, making it the first reference genome to break “usability barrier”. Having a comprehensive cannabis genome opens the door to a host of industry innovations, including marker-assisted breeding and seed-to-sale tracking systems.
Bt:Bd Allele Coverage
Modern sequencing technology can’t read the entire cannabis genome at once. Instead, the DNA is broken up into smaller more manageable pieces and then reassembled using computer software to create the full picture.
Earlier references attempted to arrange billions of small pieces, containing one hundred bases, pairs into one complete genome.
And because the cannabis genome is highly repetitive, the software mistakenly surmised that similar pieces were copies of the same piece. The Jamaican Lion reference was assembled using only pieces with millions of base pairs that can cross repeat regions, providing a more accurate picture of the genome.
The Bt:Bd allele is a perfect example of this. Early reference genomes could not differentiate between the active and inactive versions of the CBDA synthase gene. This made it very difficult, if not impossible, to determine a cultivar’s cannabinoid type. With the Jamaican Lion reference, we can see coverage over the active CBDA, inactive CBDA, and active THCA genes, and accurately call the plant type.
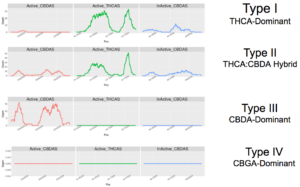
In time, we will be able to provide coverage information for other high-value genes. We are currently analyzing whole genome sequence data from 40 different cannabis and hemp cultivars with diverse cannabinoid and terpene profiles so we can better locate those genes.
Nearest Genetic Relatives and Most Genetically Distant Strains
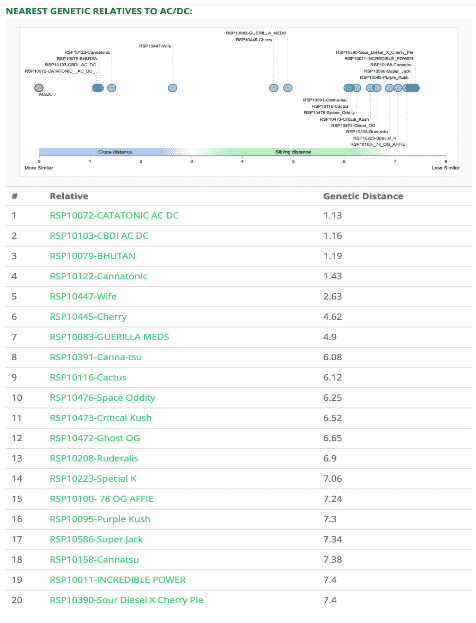
With the StrainSEEK® sequence reads mapped to the Jamaican Lion reference, we are able to more accurately determine how a cannabis cultivar compares to others in the database. That means StrainSEEK® customers may see different cultivars in their Nearest Genetic Relatives and Most Genetically Distant Strains tables. They may also notice the genetic distance values have changed.
New VCF Files and Blockchain Registration Information
Mapping to the Jamaican Lion reference generates a new VCF file, which we have submitted to the blockchain to serve as a timestamped public record of the cultivar’s genetics. Don’t worry, we have preserved the links to the original blockchain transactions along with the previous timestamp.
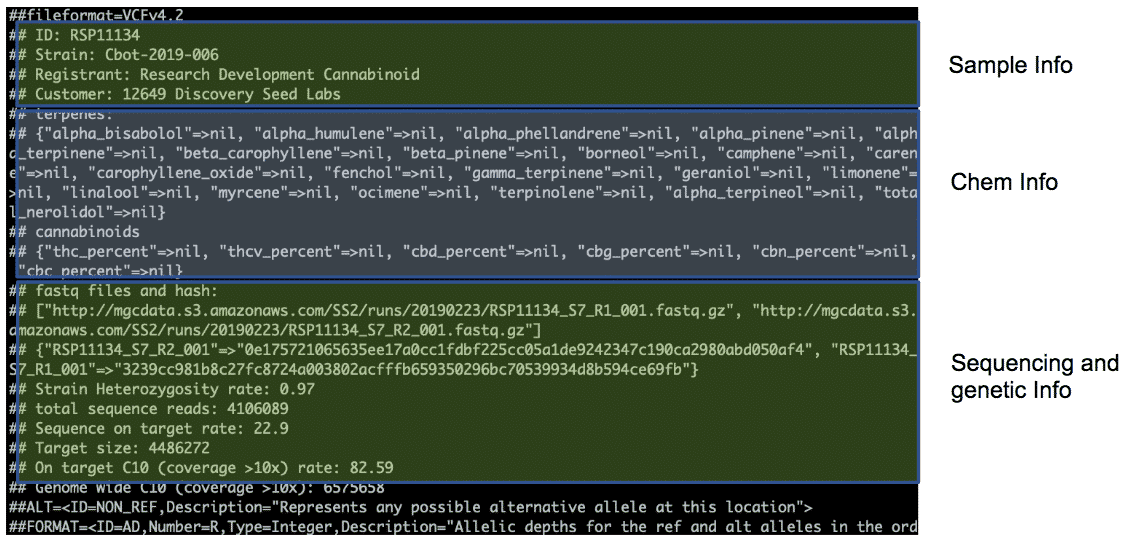
We made modifications to the VCF file to include sample information and chemotype information (when provided by the customer), as well as sequencing data summary. These modifications make the blockchain submission more powerful as proof of ownership. Meanwhile, customers can use common bioinformatics tools, such as VCFtools, GATK, and Picard to process the genetic information in the VCF file.
More Updates to Come
We are currently involved in several exciting sequencing projects that will provide additional insights into the cannabis genome that we will share with our StrainSEEK® customers.
“The annotation of structural variations in the cannabis genome will be critical to understanding the genetics of yield, seed production, and other desirable traits,” said Timothy Harkins, MGC Senior Advisor. “We continue to build new genetic tools for the entire community by using the Jamaican Lion assembly combined with the methylation and transcriptome maps and now the recombination hotspots. This combination of data and technologies are accelerating our insights, which breeders and growers can now take advantage of. None of it would be possible, however, without the extraordinarily long sequencing read lengths generated by the PacBio platform. This genetic tool wasn’t available until they came along, and that has made all the difference.”