There are currently no viable treatment options available for Hop Latent Viroid (HLVd) infected cannabis plants. Meristem tissue culture can be used to rescue infected plants, but the process can take up to 3-9 months and doesn’t always result in viroid-free plants. The success rate of tissue culture remediation is often cultivar-specific. Cultivators will surely put in the time and effort to save valuable heirloom cultivars, but many others will scrap infected plants and start a new phenohunt to replace them.
Prevention is surely the best way to protect your operation. There are a number of HLVd testing kits available that cultivators can use to confirm suspected infection or identify asymptomatic plants. Screening mother plants regularly, especially before taking a round of cuttings, would help to ensure only clean plants make it into production. Cultivators should also use extreme caution when introducing new genetics from another facility. New clones should be quarantined and screened for HLVd and other pathogens before they are allowed to share a space with existing plants.
A Case for Using qPCR for HLVd Testing
Medicinal Genomics was the first company to bring Eiken Chemical Co, Ltd.’s patented Loop-Mediated Isothermal Amplification (LAMP)-based testing to the Cannabis field with our youPCR® Gender and THC/CBD testing. We learned a lot from this endeavor and have a few thoughts to share as to why we have not yet deployed this patented technology to Hop Latent Viroid (HLVd) testing out of the gate.

First, the LAMP technology is well suited for binary questions like sex testing or the presence or absence of a given allele. For this, LAMP is fast and amenable to low cost and point-of-grow use. However, LAMP is not the best technology to address nascent biological problems like HLVd testing.
Here are a few points to consider…
Internal controls
LAMP is usually a single-plex reaction. You can amplify only one target, which makes internal controls nearly impossible. Without internal controls, you don’t know if the plant was sampled correctly. A negative result could be the result of a botched prep or it could be a true negative from a clean plant. Are you willing to bet your farm on that coin toss?
qPCR doesn’t have this problem. We can use multiple wavelengths to detect more than one target, which means we can use an internal cannabis control to ensure the DNA and RNA extraction was successful. Without internal controls, you run the risk of lots of false negatives.
This is even more problematic with RNA preps that are used for virus and viroid detection.
For example, Dadouh et al. demonstrates 1,000-10,000 fold variances in swabbing patients’ noses for RNA viruses. This paper warns against the use of a single viral Ct value in qPCR to determine viral load as you need two Ct values to properly quantitate a viral load (host and pathogen). If the sampling can vary 1,000-10,000 fold, you can easily fall off your limit of detection (LOD) and never know it. This gets even worse for +/- assays that don’t even provide a single Ct score.
Contamination Risk
Opening a tube with amplified DNA or RNA risks contaminating your facility with a PCR product that could cause false positives. LAMP’s amplification produces 10x more DNA than qPCR, making it 10x more prone to contamination. This means you should never open up a LAMP amplification tube in your facility. For this reason, we never design qPCR or LAMP assays that require the user open the tube after amplification.
Ability to Quantitate
According to MIQE guidelines, you need two bits of information in order to generate a valid viroid load: the viroid Ct score and the host Ct score. Without both, you don’t know if you are looking at a false positive, false negative, or lightly infected plant.
But why quantitate viroid load? Shouldn’t we just scrap every plant that tests positive? Maybe, but it’s difficult to cull an heirloom mother plant based on a single LAMP test when you don’t know how much viroid is a problem. Do you have 10 million copies or 10 copies? The 10 copies could be contaminated or false positives with LAMP. The 10 Million copies are more of a concern.
Given how little we know about HLVd, we believe quantitative tools are the best place to start so growers can learn if their methods are reducing viroid load or making matters worse. This can’t be done with binary, LAMP-based tests as no viroid load is provided by the test. One must resort to hyper sampling 10-50 spots on the plant to see if the percent positive changes substantially to emulate the information you get from a truly quantitative test.
Scalability
In our experience, a few dozen tests is not enough to get a full picture of HLVd contamination in a facility. You want to perform the test in duplicate to triplicate across many plants and many time points. You will find yourself in 96 well plates before you know it, so you need a platform that can manage that data torrent. qPCR platforms are ideal for this. Many LAMP-based protocols were quickly developed for SARS CoV2 testing, but few made it into high throughput production use because the CT score was too valuable to dismiss when deciding who was infectious and who was just carrying dead viral fragments.
Confidence in the call and population wide screening
One of the challenges with quantitative-blind testing at population scale is that you lose a tremendous amount of information that can be used to determine on the likelihood of the call being correct. This produces a sea of questionable data at scale. This happened in SARs-CoV-2 testing. In some labs, qPCR CT scores were calibrated to PFU/ml and suggested only calling CT <33 as cases. More broadly, tests were given FDA EUA approval without such calibration and uncalibrated CT thresholds of 37-45 were issued for many popular kits in the marketplace. When CT scores are not linked to biological infectiousness, we are forced to quarantine 5 non-infectious people for every infectious person that is PCR positive. Liotti et al. demonstrated this showing people being PCR positive for 35 days but only infectious for 7 days. In addition to the infectious versus noninfectious PCR positives, many labs were using late CTs to call positives (<37) but only using low CT samples (<32) to sequence validate their patients. When more aggressive attempts were made to sequence validate the late CT samples, many off target viruses were detected. Running this type of screening blind to CT scores is excellent for the test providers but a fool’s lottery for the grower.
Fool’s Lottery
Let’s have a look at Bayes theorem to understand this problem. Example replicated from Mathew Crawford.
This test is not unlike the conditions you will find yourself in if you test for HLVd without internal controls. Let’s assume you have a 99% chance of detecting a positive in a positive plant. You will also have some preps that fail, and, you will be unaware of this failure as you have no internal control to signal when your assay is broken as opposed to just negative. Your signal for your negative plants overlaps with your signal for bad RNA prep/PCR. This inflates False Negatives. More relevant to this example is LAMP contamination that can occur when you start opening up post amplification LAMP tubes. This can create False Positives such that a small percentage of the negatives test positive but since negatives are 98% of the data, this translates into a sizable number of all positive tests. What does this translate to when you have a low prevalence of disease (2%)?
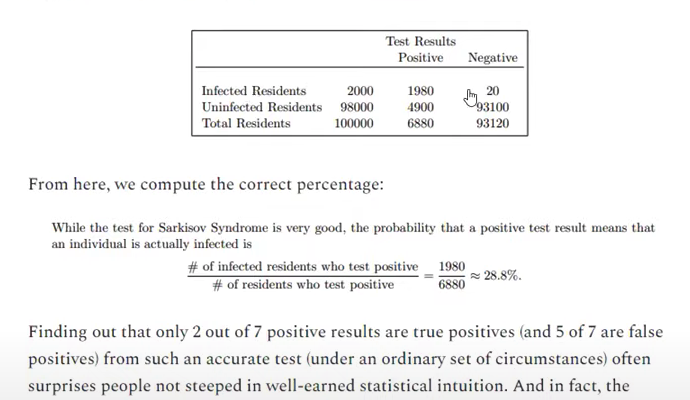

Trying to traverse this statistical mine-field with no internal controls and no CT scores is akin to gambling.
The solution to this problem is running internal controls to guarantee negatives are true negatives, and following the CTs scores of your plants across time. It should be noted that this problem is so challenging mainly because you are testing population wide. You are not focusing on only symptomatic people/plants so the vast majority of your tests will be negative. The problem with Latent viruses is their latency: They largely exist symptom-free until they don’t, so you are stuck playing the Fools Lottery unless you design calibrated methods to move the odds in your favor.
An example of qPCR HLVd testing in action
The example below shows qPCR results from HLVd infected leaves and roots. You will notice that when the plants have HLVd, they have lots of it (Ct <25) in the leaves and similar CT scores in the roots. The amount of viroid RNA (Blue) is 100X-1000X more than plant DNA (Green). There is not a tiny amount of this viroid around. When it is there, it is hot as hell in multiple diverse tissues. More sensitive tests that do not provide a quantitative value could lead you into destroying your farm over false positives.
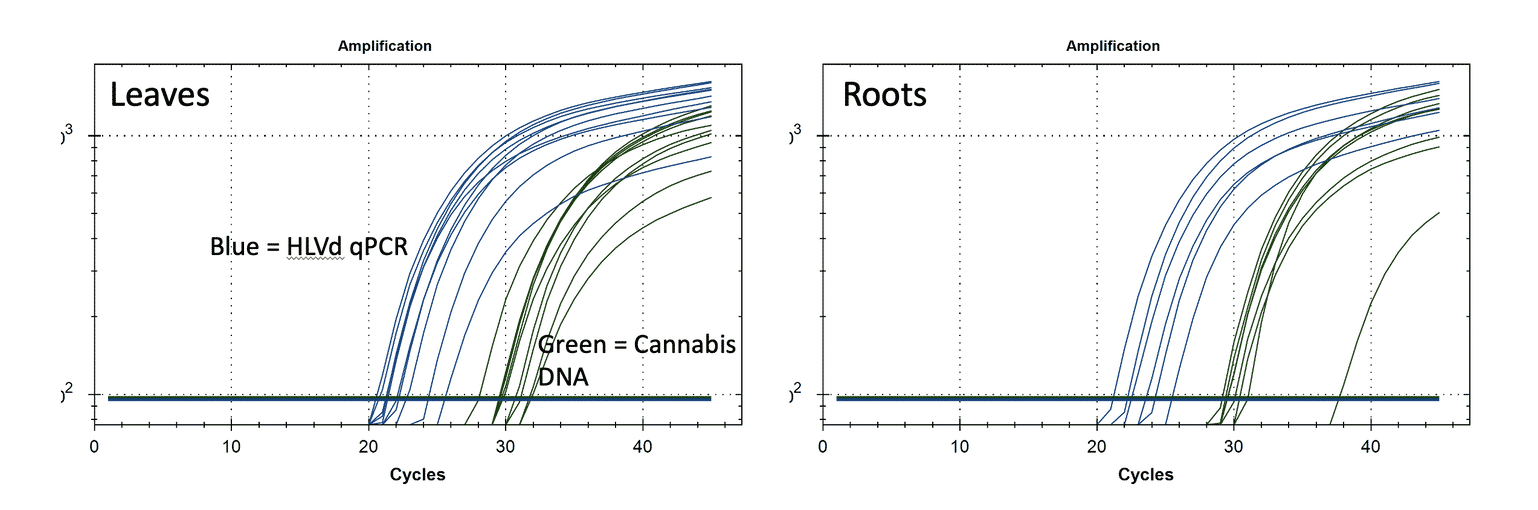
Given it is early, and nurseries are trying to learn what levels of Viroid RNA are non-clinical, or which treatments or tissue culture activities are moving the needle in the right direction, quantitative tests are far more informative, less prone to contamination, more scalable, and more actionable.
As the market matures and we learn more about the copy number of the viroid throughout its disease progression, the reproducibility of home testing and extraction will make our LAMP-based HLVd test a nice market fit. But today, we feel qualitative results will leave many customers with more questions than answers.
Regardless of when/if we roll out a LAMP-based HLVd test, we will only implement closed tube tests. The contamination problem is a sure-fire way to destroy trust in the nascent HLVd testing space. LAMP is a promising technology that is very sensitive but often this increase in sensitivity comes with trade-offs on specificity. There is no probe in LAMP that verifies your amplification event with an additional sequence so you must be very careful deploying it in population wide screens of asymptomatic targets.